Скачать с ютуб Rapid identification of bacterial invasion: full-length 16S or adaptive sampling? в хорошем качестве
Скачать бесплатно Rapid identification of bacterial invasion: full-length 16S or adaptive sampling? в качестве 4к (2к / 1080p)
У нас вы можете посмотреть бесплатно Rapid identification of bacterial invasion: full-length 16S or adaptive sampling? или скачать в максимальном доступном качестве, которое было загружено на ютуб. Для скачивания выберите вариант из формы ниже:
Загрузить музыку / рингтон Rapid identification of bacterial invasion: full-length 16S or adaptive sampling? в формате MP3:
Если кнопки скачивания не
загрузились
НАЖМИТЕ ЗДЕСЬ или обновите страницу
Если возникают проблемы со скачиванием, пожалуйста напишите в поддержку по адресу внизу
страницы.
Спасибо за использование сервиса savevideohd.ru
Rapid identification of bacterial invasion: full-length 16S or adaptive sampling?
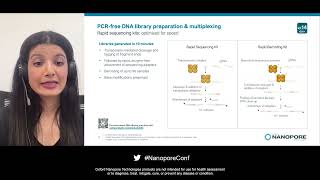
Amniotic infections can cause preterm birth and serious complications for both mother and baby. Early detection and diagnosis are crucial, but traditional methods can be slow and miss some cases. Using two different nanopore technology targeted sequencing approaches, we amplified full-length 16S rRNA and applied adaptive sampling to detect bacteria directly from clinical research specimens. The full-length 16S rRNA method enriched any bacteria that 16S primers could capture and allowed us to identify species and report polymicrobial species that the Sanger method could not. We also tried adaptive sampling to deplete host DNA and identified Prevotella sp. within 15 minutes of sequencing. In the same day, we obtained the complete genome of the bacteria and identified antimicrobial resistance genes using the sequence data. Our research demonstrates the potential use of nanopore-based bacterial identification methods to diagnose amniotic infections. Learn more about clinical research: https://nanoporetech.com/applications...